|
Thermo Fisher
neuron differentiation medium (1:1 mixture of dmem/f12 and neurobasal medium, supplemented with b-27 and n2 supplements) Neuron Differentiation Medium (1:1 Mixture Of Dmem/F12 And Neurobasal Medium, Supplemented With B 27 And N2 Supplements), supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/neuron differentiation medium (1:1 mixture of dmem/f12 and neurobasal medium, supplemented with b-27 and n2 supplements)/product/Thermo Fisher Average 90 stars, based on 1 article reviews
neuron differentiation medium (1:1 mixture of dmem/f12 and neurobasal medium, supplemented with b-27 and n2 supplements) - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
PromoCell
differentiation medium supplemental mix Differentiation Medium Supplemental Mix, supplied by PromoCell, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/differentiation medium supplemental mix/product/PromoCell Average 96 stars, based on 1 article reviews
differentiation medium supplemental mix - by Bioz Stars,
2026-04
96/100 stars
|
Buy from Supplier |
|
Millipore
dulbecco’s modified eagle’s medium (dmem Dulbecco’s Modified Eagle’s Medium (Dmem, supplied by Millipore, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/dulbecco’s modified eagle’s medium (dmem/product/Millipore Average 90 stars, based on 1 article reviews
dulbecco’s modified eagle’s medium (dmem - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Lonza
osteogenic differentiation medium Osteogenic Differentiation Medium, supplied by Lonza, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/osteogenic differentiation medium/product/Lonza Average 90 stars, based on 1 article reviews
osteogenic differentiation medium - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Cyagen Biosciences
osteogenic differentiation medium Osteogenic Differentiation Medium, supplied by Cyagen Biosciences, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/osteogenic differentiation medium/product/Cyagen Biosciences Average 90 stars, based on 1 article reviews
osteogenic differentiation medium - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
GE Healthcare
n2b27 medium N2b27 Medium, supplied by GE Healthcare, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/n2b27 medium/product/GE Healthcare Average 93 stars, based on 1 article reviews
n2b27 medium - by Bioz Stars,
2026-04
93/100 stars
|
Buy from Supplier |
|
PromoCell
skeletal muscle growth supplement mix Skeletal Muscle Growth Supplement Mix, supplied by PromoCell, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/skeletal muscle growth supplement mix/product/PromoCell Average 93 stars, based on 1 article reviews
skeletal muscle growth supplement mix - by Bioz Stars,
2026-04
93/100 stars
|
Buy from Supplier |
|
Thermo Fisher
nk cell differentiation b0 medium 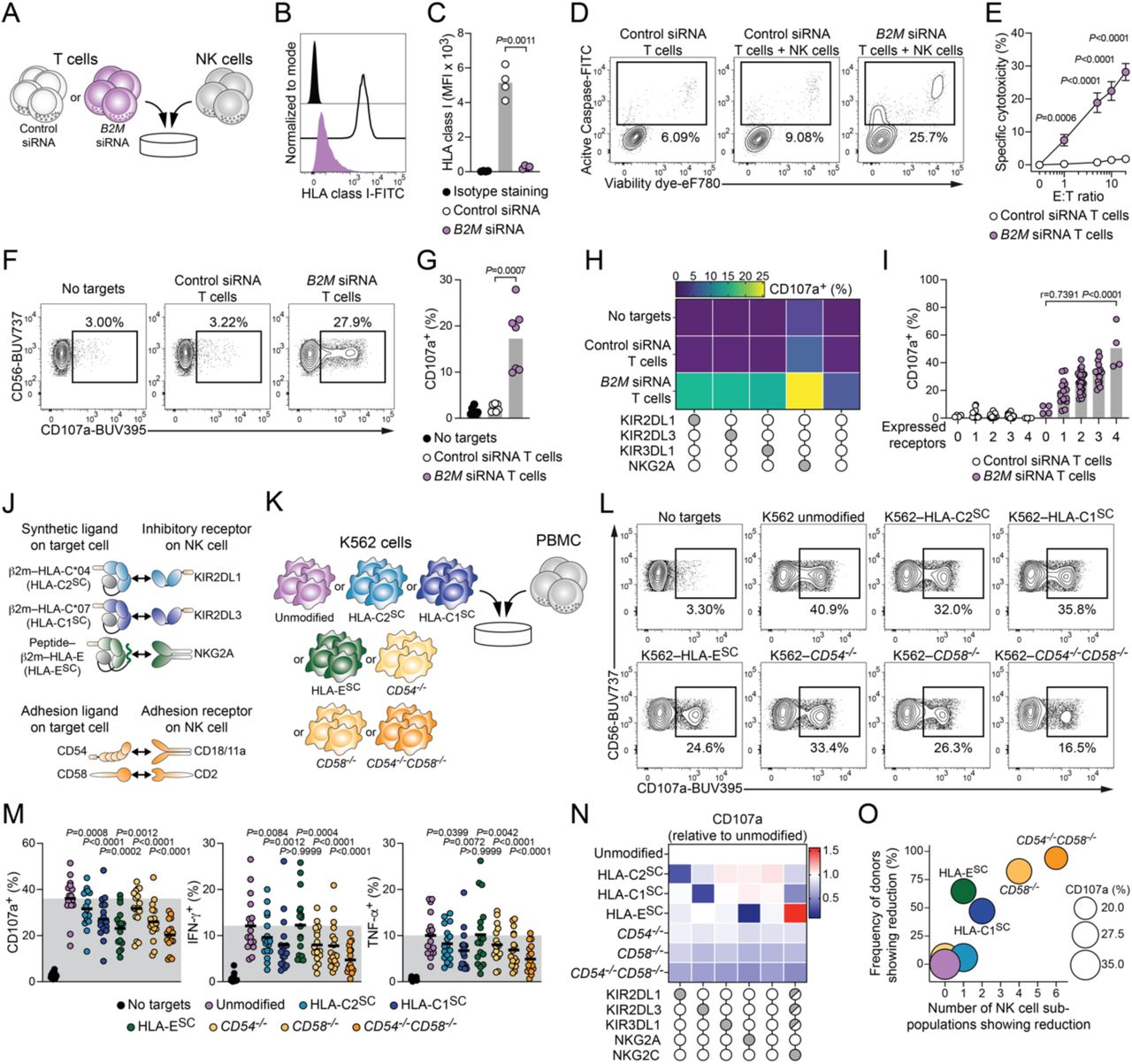 Nk Cell Differentiation B0 Medium, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/nk cell differentiation b0 medium/product/Thermo Fisher Average 99 stars, based on 1 article reviews
nk cell differentiation b0 medium - by Bioz Stars,
2026-04
99/100 stars
|
Buy from Supplier |
|
Millipore
f10 medium  F10 Medium, supplied by Millipore, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/f10 medium/product/Millipore Average 90 stars, based on 1 article reviews
f10 medium - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
TaKaRa
human cord blood cd34 hematopoietic stem cell population  Human Cord Blood Cd34 Hematopoietic Stem Cell Population, supplied by TaKaRa, used in various techniques. Bioz Stars score: 95/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/human cord blood cd34 hematopoietic stem cell population/product/TaKaRa Average 95 stars, based on 1 article reviews
human cord blood cd34 hematopoietic stem cell population - by Bioz Stars,
2026-04
95/100 stars
|
Buy from Supplier |
|
MedChemExpress
differentiation medium  Differentiation Medium, supplied by MedChemExpress, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/differentiation medium/product/MedChemExpress Average 96 stars, based on 1 article reviews
differentiation medium - by Bioz Stars,
2026-04
96/100 stars
|
Buy from Supplier |
|
CHROMagar - DRG
tm o157 differential medium  Tm O157 Differential Medium, supplied by CHROMagar - DRG, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/tm o157 differential medium/product/CHROMagar - DRG Average 90 stars, based on 1 article reviews
tm o157 differential medium - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
Image Search Results

Journal: bioRxiv
Article Title: Genetic ablation of adhesion ligands averts rejection of allogeneic immune cells
doi: 10.1101/2023.10.09.557143
Figure Lengend Snippet: (A) Schematic illustration of in vitro rejection assays between siRNA-treated T cells and NK cells from unrelated healthy donors. (B) Representative HLA class I expression on T cells following siRNA treatment. (C) Summary of HLA class I expression on T cells (n=4 donors in 4 independent experiments). (D) Representative killing of indicated siRNA-treated T cells by NK cells. (E) Summary of specific cytotoxicity (n=7 pairs in 5 independent experiments). (F) Representative degranulation of CD56 dim NK cells against indicated siRNA-treated T cells. (G) Summary of CD56 dim NK cell degranulation (n=7 donor pairs in 2 independent experiments). (H) Degranulation response pattern of CD56 dim NK cell sub-populations stratified for KIR2DL1, KIR2DL3, KIR3DL1, and NKG2A (mean of n=7 donor pairs in 2 independent experiments). (I) Degranulation of CD56 dim NK cell sub-populations stratified for number of receptors (n=4 donor pairs in 1 independent experiment). (J) Top: schematic illustration of synthetic HLA molecules as ligands for indicating inhibitory NK cell receptors. Bottom: schematic illustration of adhesion ligands and their cognate receptors. (K) Schematic illustration of in vitro NK cell activation experiments between genetically modified K562 target cells and PBMC. (L) Representative degranulation of viable CD14 − CD19 − CD3 − CD56 dim NK cells against indicated K562 target cells. (M) Summary of CD56 dim NK cell degranulation (left), IFN-γ expression (middle) and TNF-α expression (right). Grey shaded area indicates mean response against unmodified K562 target cells (n=17 donors in 5 independent experiments). (N) Degranulation response of indicated CD56 dim NK cell sub-populations against genetically modified K562 target cells relative to unmodified K562 target cells (n=17 donors in 5 independent experiments). (O) Frequency of donors that show reduction plotted against number of CD56 dim NK cell sub-populations that show reduction. Bubble size indicates degranulation of CD56 dim NK cells. Reduction is defined as relative decrease of at least 25% compared to unmodified K562 cell targets (n=17 donors in 5 independent experiments). C,G: paired t-test. E: Repeated measures two-way ANOVA with Šídák’s multiple comparisons test. I: Pearson correlation. M: Repeated measures one-way ANOVA with Šídák’s multiple comparisons test to unmodified K562 target cells.
Article Snippet: In brief, iPSC-derived CD34 + cells were transferred to
Techniques: In Vitro, Expressing, Activation Assay, Genetically Modified

Journal: bioRxiv
Article Title: Genetic ablation of adhesion ligands averts rejection of allogeneic immune cells
doi: 10.1101/2023.10.09.557143
Figure Lengend Snippet: (A) Schematic illustration of conjugation and cytotoxicity assays between K562 cells and NK cells. (B) Representative conjugate formation of CFSE-labelled NK cells to CellTrace Violet-labelled target cells. (C) Summary of conjugate formation (n=8 donors in 3 independent experiments). (D) Representative killing of target cells by NK cells. (E) Summary of specific cytotoxicity (n=8 donors in 4 independent experiments). (F) Schematic illustration of in vitro microwell assays between K562 cells and NK cells. (G) Representative display of co-cultures between single NK cells and multiple K562 cells over time. Scale bar 10 μm. (H) Summary of relative time of NK cells spent in contact with target cells (n=4 donors in 3 independent experiments). (I) Summary of relative NK cell cytotoxicity (n=4 donors in 3 independent experiments). (J) Schematic illustration of in vitro conjugation and cytotoxicity assays between mixed K562 cells and NK cells. (K) Representative deconvolution of mixed target cells based on CellTrace Violet intensity. (L) Summary of distribution of target cell type within formed conjugates (n=8 donors in 5 independent experiments). (M) Summary of distribution of target cell type within surviving target cells in cytotoxicity assays (n=8 donors in 4 independent experiments). (N) Schematic illustration of in vitro microwell assays between mixed K562 cells and NK cells. (O) Representative display of co-cultures between single NK cells and multiple mixed target cells over time. Scale bar 10 μm. (P) Summary of distribution of target cell type killed in first cytotoxic event (n=335 individual NK cells; n=4 donors in 3 independent experiments). (Q, R) Sequential killing of mixed target cells by single NK cells. (Q) Distribution of target cell types with which single NK cells form contacts but do not kill prior to killing another target cell (n=39 sparing events prior to a kill; n=4 donors in 3 independent experiments). (R) Distribution of target cell types which single NK cells kill after having spared another target cell (n=39 killing events following a spare; n=4 donors in 3 independent experiments). C, E, L, M: Repeated measures two-way ANOVA with Šídák’s multiple comparisons test. H, I: Paired t-test.
Article Snippet: In brief, iPSC-derived CD34 + cells were transferred to
Techniques: Conjugation Assay, In Vitro
Journal: bioRxiv
Article Title: Genetic ablation of adhesion ligands averts rejection of allogeneic immune cells
doi: 10.1101/2023.10.09.557143
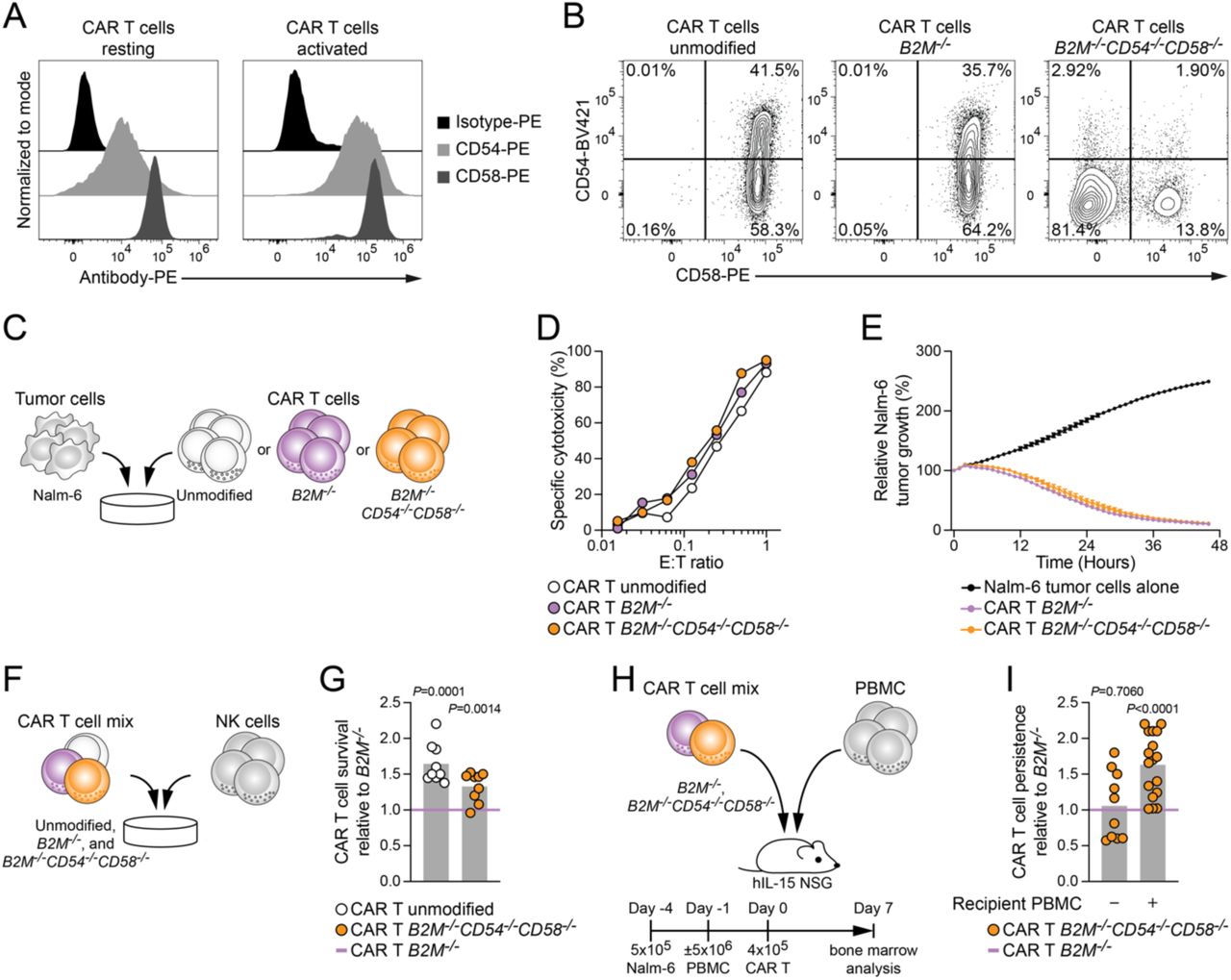
Figure Lengend Snippet: (A) Representative staining of CD54 and CD58 on resting (left) and activated (right) CAR T cells. (B) Representative staining of CD54 and CD58 on activated CAR T cells either unmodified (left), B2M -deficient (middle), or combined deletions of B2M , CD54 , and CD58 (right). (C) Schematic illustration of in vitro anti-tumor functional assay of genetically modified CAR T cells. (D) Cytotoxicity of indicated CAR T cells against Nalm-6 target cells for 18 h at varying E:T ratios. (E) Cytotoxicity of indicated CAR T cells against Nalm-6 target cells for 46 h at E:T= 1 (n=1 experiment). (F) Schematic illustration of in vitro rejection assays between CAR T cells and NK cells. (G) Summary of CAR T cell survival relative to B2M −/− CAR T cells upon co-culture with NK cells (n=9 donors in 3 independent experiments). (H) Schematic illustration of in vivo rejection assays between CAR T cells and PBMC engrafted into human IL-15-transgenic NSG mice. (I) Summary of CAR T cell persistence relative to B2M −/− CAR T cells 7 days post transfer (n=10 animals without PBMC engraftment and n=17 animals with PBMC engraftment). G, I: One-sample t-test to CAR T B2M −/− .
Article Snippet: In brief, iPSC-derived CD34 + cells were transferred to
Techniques: Staining, In Vitro, Functional Assay, Genetically Modified, Co-Culture Assay, In Vivo, Transgenic Assay

Journal: bioRxiv
Article Title: Genetic ablation of adhesion ligands averts rejection of allogeneic immune cells
doi: 10.1101/2023.10.09.557143
Figure Lengend Snippet: (A) Representative staining of HLA class I and HLA class II on viable CD45 + CD56 + iPSC-derived NK cells either expressing CD19-CAR, IL-15/IL-15Ra, and hnCD16 alone (unmodified; left), or combined with genetic deletion of B2M and CIITA (middle), or combined with genetic deletion of B2M , CIITA , CD54 , and CD58 (right). (B) Representative staining of CD54 and CD58 on viable CD45 + CD56 + iPSC-derived NK cells with indicated genotype. (C) Schematic illustration of in vitro anti-tumor functional assay of iPSC-derived NK cells. Representative (D) degranulation, (E) IFN-γ expression, (F) TNF-α expression, of viable CD45 + CD56 + iPSC-derived NK cells with indicated genotype upon co-culture with K562 tumor target cells (n=1 experiment). (G) Schematic illustration of in vitro rejection assays between iPSC-derived NK cells and NK cells. (H) Summary of specific cytotoxicity against the indicated iPSC-derived NK cells (n=4 donors in 2 independent experiments). (I) Summary of CD56 dim NK cell degranulation against the indicated iPSC-derived NK cells (n=6 donors in 2 independent experiments). (J) Degranulation response pattern of CD56 dim NK cell sub-populations stratified for KIR2DL1, KIR2DL3, KIR3DL1, and NKG2A (mean of n=6 donor pairs in 2 independent experiments). (K) Schematic illustration of in vivo rejection assays between iPSC-derived NK cells and activated NK cells engrafted into human IL-15-transgenic NSG mice. (O) Summary of iPSC-derived NK cell persistence relative to animals without recipient NK cell engraftment (n=9-10 animals per group in 2 independent experiments). (P) Schematic illustration of rejection of B2M −/− allogeneic cells by recipient NK cells (top) and averting of rejection by combining B2M −/− with deletion of CD54 and CD58. H, I: Repeated measures two-way ANOVA with Šídák’s multiple comparisons test. L: Ordinary one-way ANOVA with Šídák’s multiple comparisons test.
Article Snippet: In brief, iPSC-derived CD34 + cells were transferred to
Techniques: Staining, Derivative Assay, Expressing, In Vitro, Functional Assay, Co-Culture Assay, In Vivo, Transgenic Assay

Journal: Iranian Journal of Basic Medical Sciences
Article Title: Synergistic bactericidal activity of a naturally isolated phage and ampicillin against urinary tract infecting Escherichia coli O157
doi: 10.22038/IJBMS.2019.37561.8989
Figure Lengend Snippet: Inhibition zones on the Escherichia coli O157 lawn. (a) the left spot is pertaining to purified phage from single plaque and (b) the right spot was produced by the phage cocktail enriched from sewage. Determination of plaque forming unit by serial dilution of the phage and evaluation of clear zones produced by spot test on TTC mixed overlay agar plates, (c) plaque forming units of phage at MOI of 10, (d) spotting serial dilution of the phage (10 µl ) on TTC mixed bacterial lawn, (e) spot test of concentrated phage
Article Snippet: This strain was verified using
Techniques: Inhibition, Purification, Produced, Serial Dilution, Spot Test

Journal: Iranian Journal of Basic Medical Sciences
Article Title: Synergistic bactericidal activity of a naturally isolated phage and ampicillin against urinary tract infecting Escherichia coli O157
doi: 10.22038/IJBMS.2019.37561.8989
Figure Lengend Snippet: Synergy between the phage and ampicillin in the endpoint microdilution assay. The percentage of the growth of Escherichia coli O157 in the presence of ampicillin at concentrations of 1.562, 3.125, 6.25, 12.5, 25 µg/ml and by the same concentrations mixed with the phage at MOI 10; the incubation time was 6 hours at 37 ° C
Article Snippet: This strain was verified using
Techniques: Microdilution Assay, Incubation

Journal: Iranian Journal of Basic Medical Sciences
Article Title: Synergistic bactericidal activity of a naturally isolated phage and ampicillin against urinary tract infecting Escherichia coli O157
doi: 10.22038/IJBMS.2019.37561.8989
Figure Lengend Snippet: Effect of the synergistic antibacterial activity of the phage and ampicillin on the Escherichia coli O157 growth curve
Article Snippet: This strain was verified using
Techniques: Activity Assay